✂️ & 🏷️ R-loop
Intro
CUT&Tag Resource Center
CUT&Tag-IT® R-loop Service
CUT&Tag-IT® R-loop Assay Kit
CUT&Tag-IT® Spike-In Control, R-loop
1. Cut & Tag protocol
Note
we used reduced volumes of reagents/solvents when we weren’t sure if we had enough (specified in Marketa’s protocol)
PDF: Protocol; CUT&Tag-IT™ R-loop Assay Kit
PDF: CUT&Tag-IT® Spike-In Control, R-loop
PDF: Counting Cells and Nuclei for Epigenetic Applications
Library Preparation for Illumina
- I find this one very usefull
- it includes also videos with “real life tips” on right side - description of each bellow
Library Preparation for Illumina: text & videos
2. Concentration
Qubit & Tapestation
For a professional NGS library prep, the standard protocol:
Use the TapeStation for the “Size” and the Qubit for the “Concentration.”
- When pooling libraries for an Illumina run, even a small error in concentration can lead to “index hopping” or uneven data distribution (where one sample gets 90% of the reads and others get 10%).
- Using the TapeStation to check size and the Qubit to check molarity ensures your final calculations for pooling (converting ng/µL to nM) are as accurate as possible.
When can you skip one?
If you are in a situation where you must choose just one to save time or sample:- TapeStation only : samples are all the exact same type (e.g., all the same PCR amplicon length) and you have a very high concentration.
- Qubit only: already know exactly what size your fragments are (e.g., a specific 300bp PCR product) and don’t need to check for degradation.
(source: Gemini)
2A. Qubit
Note
- protocols are usualy everywhere with Qubit
- there are special tubes for it
Qubit dsDNA Quantification Assay Kits- dsDNA
- dsDNA: High Sensitivity
- lower and upper point calibration (S1 and S2)
- dilution of samples 1:200 : 1 μL + 199 μL
- units: ng/μL
Workflow
- samples on ice
- thaw at r.t. (but keep on the ice rest of the time)
- “flick the samples” – to homogenize
- spin down (short spin at r.t. centrifuge)
- mix with buffer (pipet first buffer 199μL and then sample 1μL)
- quick vortex
- incubation (r.t.; 3 min; can differ base of kit)
- measure (ng/μL) + write down a concentrations
- optional: download data on USB
Qubit™ 4 Fluorometer: pg 18-28
Calculation of dsDNA library concentration
- if you would like to calculate concentration from Qubit
- dsDNA library concentration
2B. Tapestation
- Agilent High Sensitivity D1000 ScreenTape Assay Quick Guide for 4200 TapeStation System
- You Toube: TapeStation tutorial - very nicely explained; Workflow: sapmple preparation from 01:00; Workflow: TapeStation from 08:27; Workflow: Software from 09:37
- You Toube: Agilent TapeStation: DNA and RNA Applications; Workflow from 14:54
- Agilent TapeStation
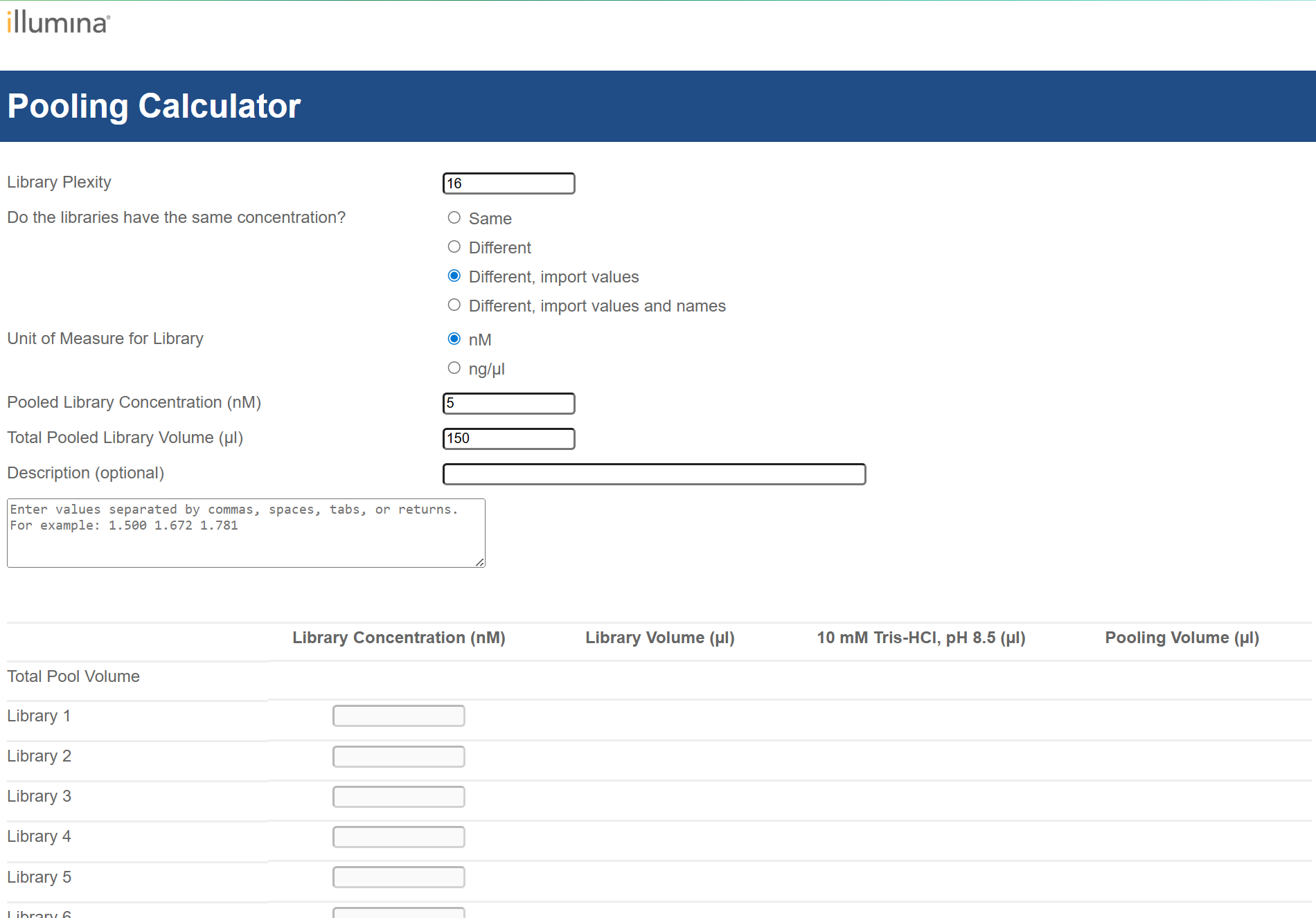
3. Pooling
- concentration from libraries
- elution buffer as diluent for pooling - correct concentration of Tris
Novaseq X
- pool samples equimolarly
- the minimum 5nM to guarantee the best results
- 35 μL per lane (minimum volume); can be higher
- pool in a 1.5 mL low-bind tube
Under-represented samples
Example
- pool all experimental samples equimolarly (5nM)
- pool the 4 negative controls at a slightly lower molarity
It is better to discuss dilution with the genomics center before pooling.
Let them know, which assay was used at TapeStation - for their quality assesment (HS D1000, or they can try D5000 for longer fragments).
Further resources
NEBNext® Ultra™ II DNA Library Prep Kit for Illumina®
Multiplex Oligos for Illumina Protocol
Learn about Illumina’s Next-Generation Sequencing Workflow
Illumina’s Next-Generation Sequencing Workflow